Database model¶
Database model was built based on Project Workflow (see: Project Workflow Overview).
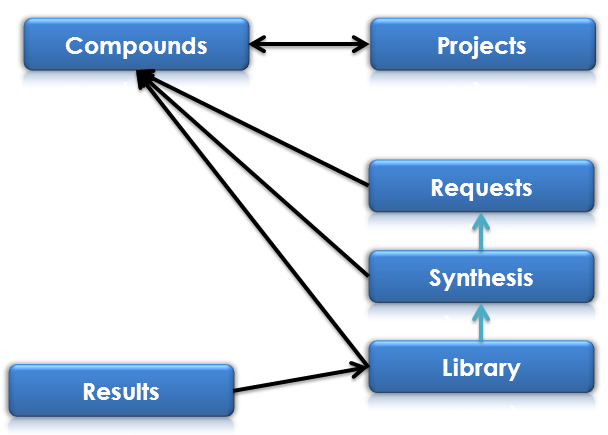
Simple Model¶
Compounds and Projects tables are connected by many to many relationship with allowing to adding one compound to many projects and many compounds to one project.
Compounds Table is containing basic information about molecule struture like:
- SMILES and InChi code,
- name
- fingerprints
- molecular weight
- logP
- numer of atoms with and without Hydrogens
- number of rings
- Hbond acceptors
- Hbond donors
- Teorethical PSA
Requests, Synthesis and Library tables are refering to this data by relationship to the Compounds. This mean that editing and changing structure for Compound will make changes for all branches in all connected tables.
Synthesis table is storing ID of Request instance from which it is derived. This making the posibility for tracking the pathway of compound from one to other table and the ability to implement The Project Workflow in a consistent way.
The same connection is between Synthesis and Library tables.
Results table has many to one relationship to Library. In this way it’s aviailble to add many results to one library instance.
Basic schema of database model is presented below:
Detailed Model¶
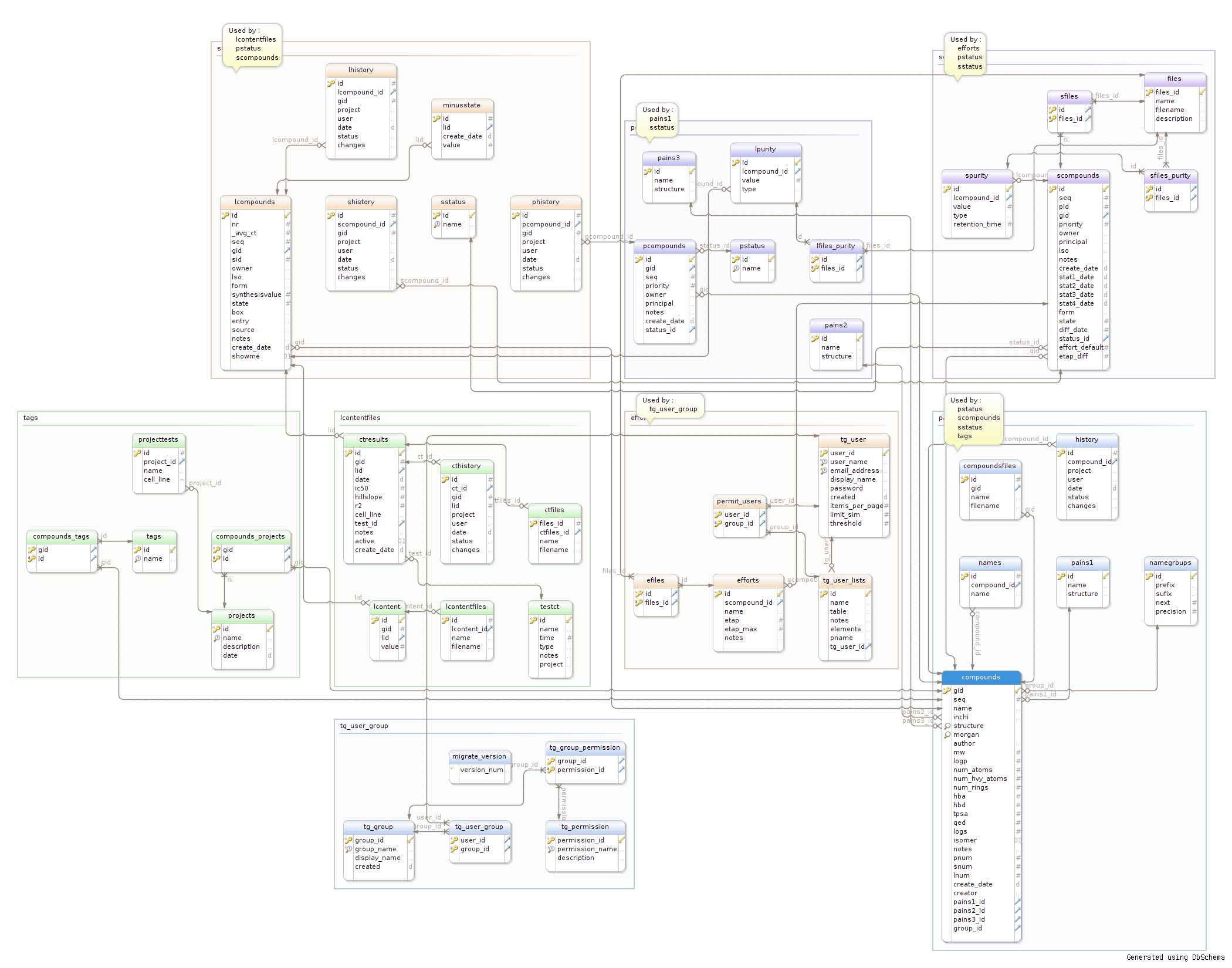
Entity - Relationship Diagram for MolGears Database model is presented below.
Diagram was generated using DBSchema.
Main 6 Tables are respectively named as:
| Model Name | SQL name |
|---|---|
| Projects | projects |
| Compounds | compounds |
| Requests | pcompounds |
| Synthesis | scompounds |
| Library | lcompounds |
| Results | ctoxicity |
- Each of the main tables (except the Projects Table) has dedicated history tables for storing changes.
- Request and Synthesis tables have dedicated tables for Status.
- Compounds, Synthesis, Library and Results tables have dedicated tables for storing files.
- There are 3 PAINS tables for storing SMARTS codes.
- The tables for access managing have “tg_” prefix (tables for Users accounts, Users Lists, Groups and Permissions).
- Other tables are auxiliary and servs i.a. for auto-naming, state and purity monitoring, test description, tags storing etc.
ER Diagram (click the image to enlarge):